Goal:
To develop genetics-related techniques, especially those essential for maintenance and supply of laboratory mice and stem cell lines at a high quality in BioResource Research Center (BRC).
Activities:
- Development of mouse somatic nuclear transfer techniques
- Development of microinsemination techniques
- Development of reliable cryopreservation techniques for mouse embryos or gametes
- Development of new stem cell lines and animal models
Specific Aims
1. Development of mouse somatic cell nuclear transfer (SCNT) techniques
Development of mouse somatic cell nuclear transfer (SCNT) techniques We have previously found that a group of paternally expressed imprinted genes were dysregulated (biallelically expressed) in mouse SCNT placentas, where an abnormal hyperplasia is consistently observed (Okae et al., Hum Mol Genet, 2014; Hirose et al., Epigenetics, 2018; Matoba et al., Cell Stem Cell, 2018). To elucidate the function of these imprinted genes in mouse placentas, we have generated following two knockout (KO) mice lines.
● Slc38a4
Slc38a4 is the gene encoding an amino acid transporter. Interestingly, paternal KO mice of Slc38a4 showed severe placental hypoplasia
(30-40% reduction in weight) associated with intrauterine growth restriction of fetuses. However, the SCNT mice generated from the donor cells
derived from maternal KO of Slc38a4 still showed abnormal placental hyperplasia (Matoba et al., PNAS, 2019). These results suggest that the
dysregulated imprinted genes other than Slc38a4 are likely responsible for the abnormal placental phenotypes in mouse SCNT embryos.
● Sfmbt2 microRNA (miRNA)
Sfmbt2 miRNA is the largest miRNA cluster in mice located in the 10th intron of Sfmbt2 gene that is essential for placental development in mice. Using the entire Sfmbt2 miRNA lacking mouse line, we demonstrated that Sfmbt2 miRNA cluster positively regulated development of the
spongiotrophoblast (ST) layer in mouse placentas (Inoue et al., Cell Rep., 2017). When we used donor somatic cells derived from the maternal KO of Sfmbt2 miRNA, the weight of SCNT placentas was significantly decreased and proliferation of the ST layers was also ameliorated (Inoue et al., in revision).
2. Development of microinsemination techniques
Although the primary spermatocyte nucleus can support full term development following injection into immature oocytes,
the birth rates are extremely low (around 4%)(Ogura et al., PNAS, 1998; Miki et al., J Reprod Dev, 2006). At least one of the causes of this low efficiency is chromosome segregation errors during
meiosis within oocytes. Then we used the “down-sized oocytes” which are known to increase the precise chromosome segregation for
injection with primary spermatocytes. As a result, the incidence of aneuploidy was significantly decreased (Table and Figure) and the birth rate was
improved to 20-30% per embryos transferred. (Collaboration with Dr. T. Hino, Asahikawa Medical University).
Table: Chromosome analysis of mouse oocyte injected with primary spermatocyte
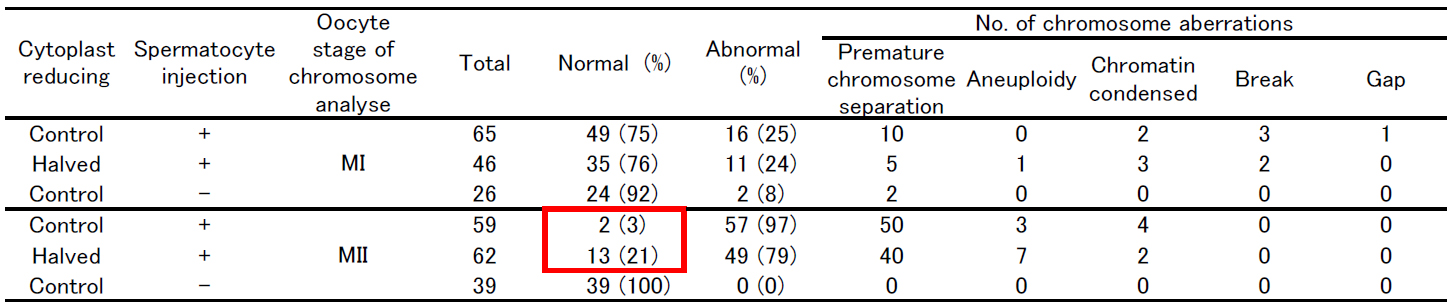
Figure: Chromosome analysis of halved oocyte injected with primary spermatocyte.
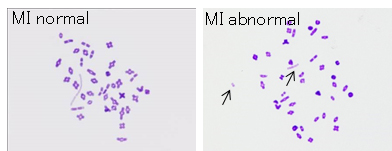

3. Development of reliable cryopreservation techniques for mouse embryos or gametes
We produced interspecies chimeric mice using ES cell lines from wild-derived mouse of ZBN/Ms and SPI/TUA (M. m. spicilegus) strans by injection into 8-cell embryos or blastocysts of ICR mice. We are confirming the germline transmission by natural mating for preservation of thesie strains by ES cells.
![Fig.1:Interspecies chemera of ES cells established form ABN/Ms(A,B)abd SPI/TUA(C,D)strain [M. Spicilegus] and blastocyst of ICR strain [M. m. domesticus]](https://kougaku.brc.riken.jp/en/wp-content/uploads/rimg/hasegawaE1.jpg)
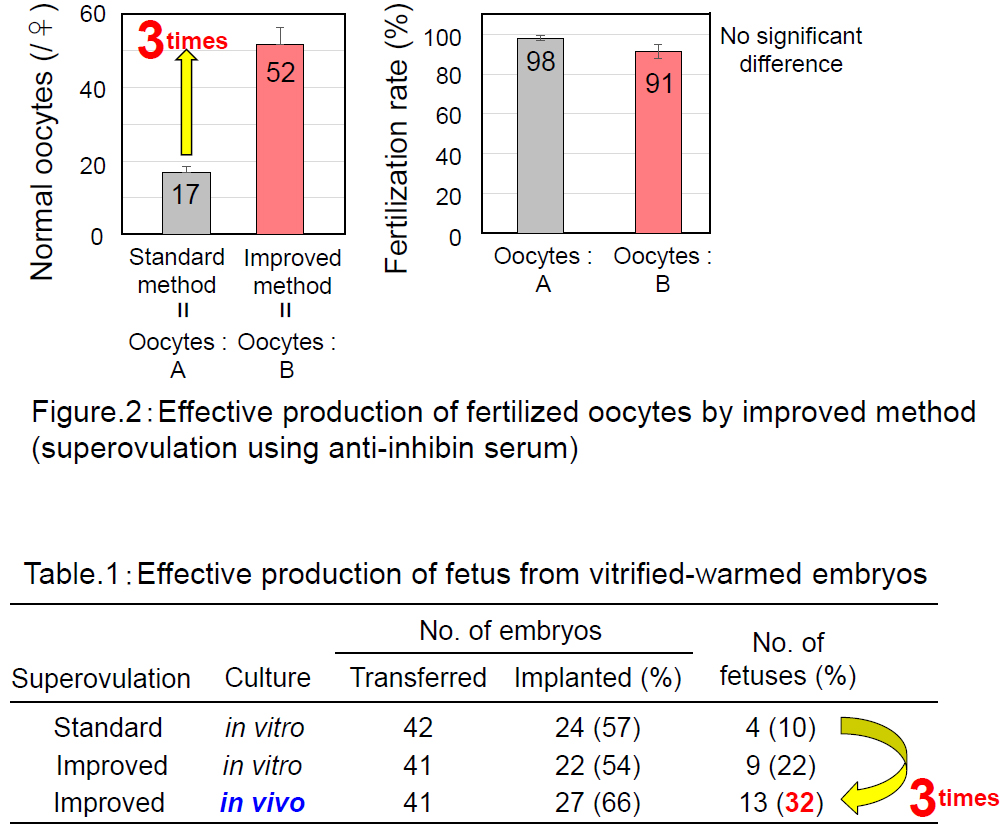
Using the “A” strain of mice that is resistant to assisted reproductive technologies, we successfully increased the number of ovulated oocytes from 17 to 52 by combining progesterone-induced estrous synchronization with anti-inhibin serum-induced superovulation (Hasegawa et al. Biol. Reprod, 2016). We could retrieve their 2-cell embryos for cryopreservation by oviductal transfer immediately after IVF. With these techniques, we increased the number of A strain pups obtained per female from 1.7 to 15.1.
4. Development of new stem cell lines and animal models
Trophoblast stem cells (TSCs) that are established in vitro from trophectoderm of blastocysts provide the best model for molecular and functional analyses of the trophoblast lineages. We have previously shown that TSCs consist of heterogeneous cell
populations (Motomura et al., Biol Reprod, 2016), suggesting that it is important to identify key factors required for TSC maintenance.
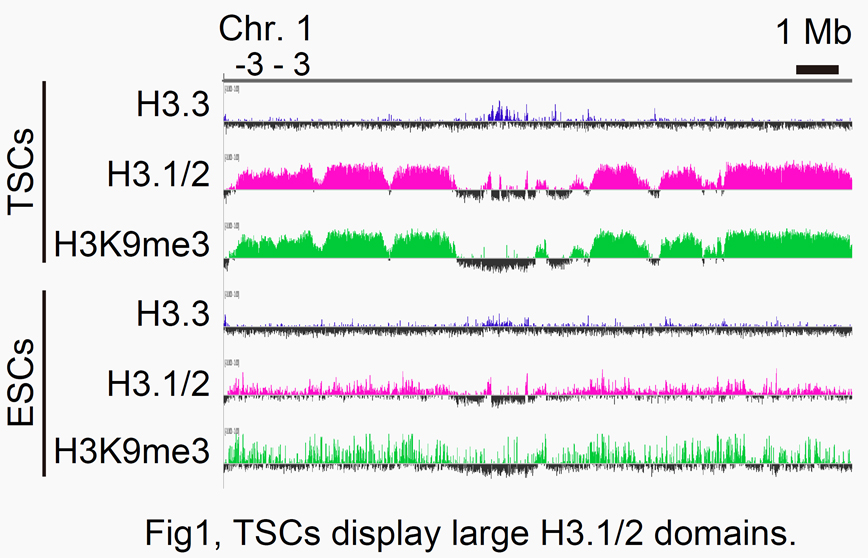
To this end, we are now focusing on the involvement of histone variants, especially H3.1 and H3.2 (H3.1/2), in the epigenome maintenance of TSCs. Our ChIP-seq analysis revealed that the TSC genome contains large H3.1/2 domains associated with H3K9me3,
a heterochromatin mark (Fig. 1). To see the biological significance of these H3.1/2 domains within the TSC genome, we knocked down P150, a major component of CAF-1 chaperone, in cultured TSCs. As a result, the expression levels of several TSC marker genes decreased,
while the expression of the ESC marker Oct3/4 increased. These data suggest that H3.1/2 is one of the key factors that determine the epigenetic nature of TSCs.
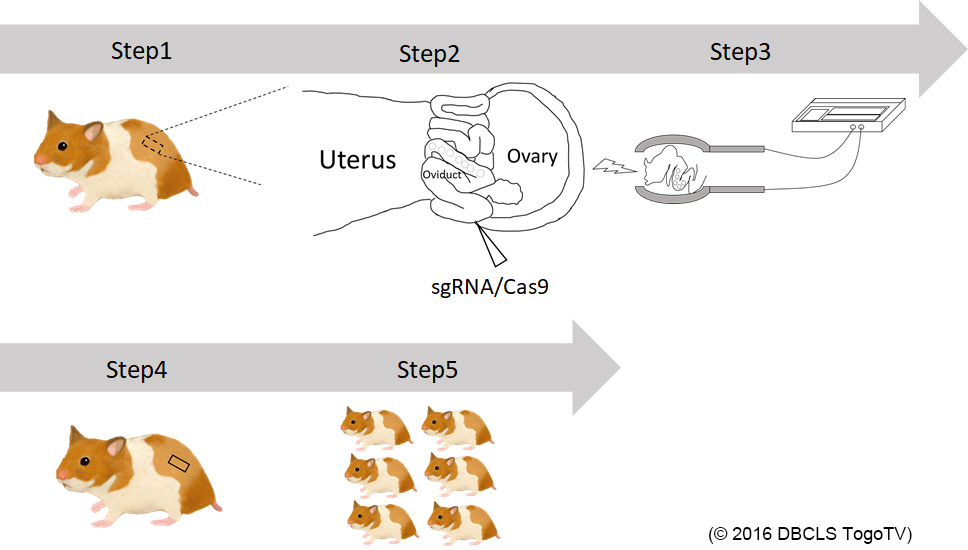
We are generating knockout hamsters for identification of the gene functions that are unknown in mouse studies (Hirose et al., PNAS, 2020). This year, we successfully generated hamsters deficient for Kir6.2, a major subunit of the ATP-sensitive K channel. We will see the glucose metabolism of homozygous knockout offspring.